To determine whether interplay between RUNX1 and FOXL2 occurs at the chromatin level, we performed genome-wide analysis of RUNX1 chromatin occupancy in E14.5 ovaries. RUNX1 plays complementary/redundant roles with FOXL2 to maintain fetal granulosa cell identity, and combined loss of RUNX1 and FOXL2 results in masculinization of the fetal ovaries. In the mouse, RUNX1 marks the supporting cell lineage and becomes granulosa cell-specific as the gonads differentiate. We discovered that the transcription factor RUNX1 is enriched in the fetal ovary in rainbow trout, turtle, mouse, and human. This process of fate specification hinges on a balance of transcriptional control. Sex determination of the gonads begins with fate specification of gonadal supporting cells into either ovarian granulosa cells or testicular Sertoli cells. Nicol B, Grimm SA, Chalmel F, Lecluze E et al. RUNX1 maintains the identity of the fetal ovary through an interplay with FOXL2
EXERCISE 3: QUESTIONS AND STEPS TO FOLLOW.EXERCISE 2: QUESTIONS AND STEPS TO FOLLOW.EXERCISE 1: QUESTIONS AND STEPS TO FOLLOW.Module 6 Lab: GeneMANIA (Cytoscape version).Create a metatargetome using iRegulon and merge 2 networks in Cytoscape. Module 5 Lab 2: Gene Regulation and Motif Analysis Practical Lab / iRegulon.Exercise 6 (optional): Get the iRegulon RUNX1 targets and find the mouse orthologs using g:Orth (from g:Profiler) to create the gmt file used in Exercise 4.
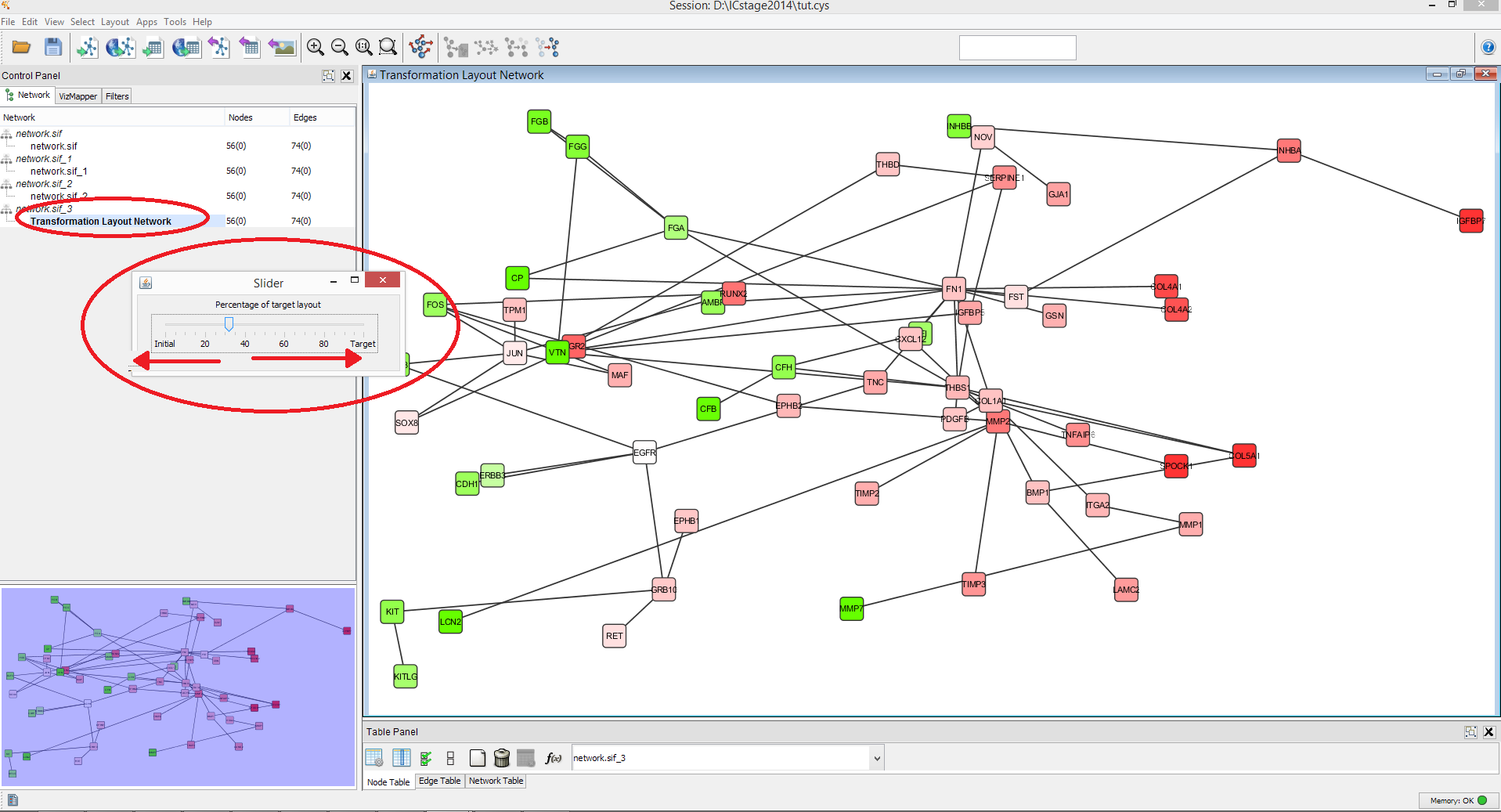
ORTHOLOG LAYOUT IN CYTOSCAPE HOW TO
Exercise 5: Learning how to run MEME-chip from the MEME suite ( ).Step 4b Optional: Change the edge style of the signature gene-sets:.Exercise 4: Add RUNX1 targets and RUNX1 KO genes on the distal enrichment map.Optional exercise 3c: Repeat the process by building both the Proximal and Distal enrichment maps at the same time.Optional exercise 3b: Repeat the process of building an enrichment map using the proximal data (Proximal_GOBP_greatExportAll.tsv).Optional exercise 3a: AutoAnnotate the enrichment map:.Exercise 3 (optional): Practice building enrichment maps and auto-annotation.Exercise 2 - Build an enrichment map to visualize GREAT results.Perform pathway enrichment - Proximal approach.Exercise 1 - Run pathway analysis using GREAT.Module 5 Lab 1: Gene Regulation and Motif Analysis Practical Lab /chIP-seq.Additional slides about the tools Segway and BEHST presented during the lecture.Practical lab 2: gene list - iREgulon and enrichr/EnrichmentMap.Practical lab 1: chIP_seq data - GREAT and MEME-chIP.Module 4: In-depth Analysis of Networks and Pathways.

Step 5 - Step through notebook to run the analysis.Create your first notebook using Docker.Set Up - Option 2 - Docker image with R/Rstudio.Exercise 4 (Optional) - Explore results in GeneMANIA or STRING.Exercise 2 - Post analysis (add drug target gene-sets to the network).Exercise 1 - GSEA output and EnrichmentMap.Exercise 1c (optional) - investigate individual pathways in GeneMANIA or String.Exercise 1c - create EM from results using Baderlab genesets.Exercise 1b - Is specifying the gmt file important?.Exercise 1a - compare different gprofiler geneset size results.Module 3: Network Visualization and Analysis with Cytoscape.Module 2: Finding Over-represented Pathways (Veronique Voisin).Module 1 - Introduction to Pathway and Network Analysis (Gary Bader).Pre-Workshop Materials and Laptop Setup Instructions.Pathway and Network Analysis of -Omics Data ( May 2021 ).
